Abstract
Introduction: The purpose of this study was to detect ybtS, entB, mrkD, magA, kfu, iutA, rmpA and K2 genes in extended-spectrum beta-lactamase (ESBL) - and non-ESBL producing Klebsiella pneumoniae.
Methods: To this end, 70 K. pneumoniae isolates were selected from hospitals of Kurdistan Province, Iran. The ESBL phenotype was conducted utilizing the disc diffusion technique in accordance with CLSI procedures. Detection of virulence factor genes was performed by the PCR in the ESBL and non-ESBL isolates.
Results: Sixty-two (88.6%) isolates of K. pneumoniae were ESBL producers. Further, entB had the most frequency in all the isolates. There were no significant differences between ESBL production and the presence of ybt S, entB, mrkD, magA, kfu, iutA, rmpA and K2 genes and the presence of these genes and variables such as presence of sex, clinical specimen type, and hvKP phenotype among the ESBL and non-ESBL K. pneumoniae isolates.
Conclusion: In conclusion, in other studies, K. pneumoniae strains were separated from liver abscesses and the magA gene was frequently present; however, in our study, the K. pneumoniae strains were separated from various clinical specimens and the magA gene had low frequency.
Background
Klebsiella pneumoniae is a prominent opportunistic pathogen which causes upper respiratory tract infection, diarrhea, pneumonia, urinary tract infection (UTI), and septicemia 123. The prevalence of drug resistance in K. pneumoniae has increased, which is because of extended-spectrum beta-lactamase (ESBL) enzymes and appearance of multi-drug resistant (MDR) K. pneumoniae 45. In addition, K. pneumoniae possesses different virulence factors that contribute to its pathogenicity including lipopolysaccharide (LPS) O-side chain (endotoxin), capsular polysaccharide, adhesions and sidrophores 647. The LPS contains lipid A, core, and O-polysaccharide antigen 8. Capsule polysaccharide (CPS) is a major factor for virulence of K. pneumoniae and classified into 77 serological types (K) 9. Capsular layers engulf the surface of bacteria and prevent bacteria phagocytosis. K1 and K2 capsular antigens are the most important ones 10.
Genome of the K. pneumoniae capsule comprises gene clusters cps (capsular polysaccharide synthesis), magA (mucoviscosity associated gene A), rmpA and wb (O-specific polysaccharide directed by the wb gene cluster) 11. MagA (35-Kbp) was identified as a K1-specific capsular polymerase gene which acts as a trans-acting activator for biosynthesis of cps. Moreover, magA is homologous with the genes involved in glycosylation, transfer and biosynthesis of the LPS12. In 2004, magA was determined as the major virulence factor of K. pneumoniae 2. It has been reported that rmpA can magnify the colony mucoidy of different serotypes of K. pneumoniae and act as a plasmid-mediated regulator of extra capsular polysaccharide synthesis 13. Adhesives include Pilli, the building of protein, and the attachment of bacteria to the host. MrkD gene mediates binding to the extracellular matrix, and also codes type 3 fimbria adhesion 14. K. pneumoniae by different siderophores (iron-bound) including enterobactin, yersiniabactin and hydroxamate siderophore obtain iron from transferrin and lactoferrin in host transport proteins. EntB, ybtS, kfu and iutA genes encode enterobactin, yersiniabactin, iron-uptake system and hydroxamate siderophore 15. The main purpose of the current study was to detect ybtS, entB, mrkD, magA, kfu, iutA, rmpA, and K2 genes in ESBL and non-ESBL producing K. pneumoniae isolated from clinical specimens in Kurdistan Province, Iran.
Methods
Identification of bacterial strains
Seventy K. pneumoniae isolates were taken from specimens including urine, blood, tracheal aspirates and wound from October 2015 to July 2016 from general hospitals of Kurdistan Province, Iran. All the isolates were cultured on blood and MacConkey agar (Merck, Germany). Colonies were identified by Gram stain and biochemical tests such as urea hydrolysis, H2S production, lysine decarboxylase, lactose fermentation, indole, methyl red, voges proskauer, citrate (IMViC) and oxidase tests 16.
Phenotypic detection of ESBLs strains
Detection of ESBLs was tested by the combination disk diffusion test (CDDT) for the K. pneumoniae isolates. The CDDT was performed by ceftazidime (30 μg), cefotaxime (30 μg), cefepime (30 μg), cefpodoxime (30 μg), cefotaxime-clavulanic acid (30/10μg), cefotaxime-clavulanic acid (30/10 μg), cefepime-clavulanic acid (30/10 μg) and cefpodoxime-clavulanic acid (30/10 μg) (Roscoe, Denmark) (Ho et al., 1998). Escherichia coli ATCC 25922 and K. pneumoniae ATCC 7881 were utilized for negative and positive controls, respectively.
Determination of hypermucoviscosity K. pneumoniae (hv-KP) phenotype
Seventy K. pneumoniae isolates were separated from the clinical samples and cultivated on blood agar medium (Merck, Germany); then, they were incubated at 37°C for 24 h. Subsequently, the hypermucoviscosity of K. pneumoniae (hv-KP) phenotype was determined by forming a viscous string more than 5 in standard bacteriological loops 3.
Virulence genes identification by PCR amplification
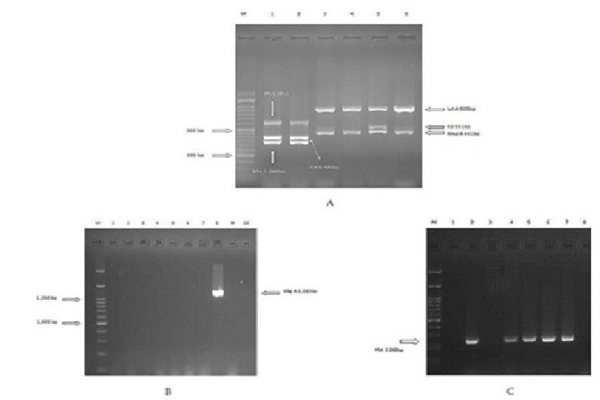
The K. pneumoniae isolates were cultured on Luria broth (LB) medium overnight. Then, DNA samples were extracted using Genomic DNA Extraction Kit (SinaClon, Iran). Gene coding virulence factors were detected by the PCR method. PCRs were carried out by using the thermo cycler system (Bio-Rad, Australia) and master mix PCR (YT1553, Iran) and primers were designed by Compain et al. (Table 1) 14. Amplification was carried out as follows: initial denaturation at 95°C for 15 min followed by 30 cycles of denaturation at 94°C for 30 s, 60°C for 90 s, and 72°C for 60 s and elongation at 72°C for 10 min. One multiplex PCR was not performed for detection of genes due to unavailability of one control isolate with multiple genes. Control positive isolates were obtained from the Lorestan University of Medical Sciences, Iran.
| Primer | Name DNA sequence (5 to 3) | Amplicon size (bp) |
| ybtS_forybtS_rev | GACGGAAACAGCACGGTAAAGAGCATAATAAGGCGAAAGA | 242 |
| mrkD_formrkD_rev | AAGCTATCGCTGTACTTCCGGCAGGCGTTGGCGCTCAGATAGG | 340 |
| entB_forentB_rev | GTCAACTGGGCCTTTGAGCCGTCTATGGGCGTAAACGCCGGTGAT | 400 |
| rmpA_forrmpA_rev | CATAAGAGTATTGGTTGACAGCTTGCATGAGCCATCTTTCA | 461 |
| K2_forK2_rev | CAACCATGGTGGTCGATTAGTGGTAGCCATATCCCTTTGG | 531 |
| kfu_forkfu_rev | GGCCTTTGTCCAGAGCTACGGGGTCTGGCGCAGAGTATGC | 638 |
| iutA_foriutA_rev | GGGAAAGGCTTCTCTGCCATTTATTCGCCACCACGCTCTT | 920 |
| magA_formagA_rev | GGTGCTCTTTACATCATTGCGCAATGGCCATTTGCGTTAG | 1283 |
Statistical analysis
The association between the ESBL production, clinical specimen type, sex ,and hvKP phenotype and presence of ybtS, entB, mrkD, magA, kfu, iutA, rmpA and K2 genes among the ESBL and non-ESBL K.pneumoniae isolates was analyzed by Fisher tests with STATA software program v12.
Results
Bacterial isolates
Out of the 70 K. pneumoniae isolates, 37 (52.9%), 32 (45.7 %) and 1 isolates (1.4%) were collected from women, men and the hospital environment. Moreover, 50 isolates (71.4%) were obtained from urine, 8 isolates (11.4%) from blood, 10 isolates (14.3%) from tracheal aspirates, 1 isolate (1.4%) from wound, and 1 isolate (1.4%) from the environment (Table 2).
| Sample No | Origin | Sex | ESBL production | Virulence genes |
| Kp6 | urine | male | positive | ybt S,entB, mrkD |
| Kp12 | urine | female | positive | ybt S,entB, mrkD |
| Kp16 | urine | male | positive | entB, mrkD, kfu |
| Kp32 | urine | female | positive | ybt S,entB, mrkD, magA |
| Kp39 | tracheal | male | positive | entB, mrkD |
| Kp43 | urine | female | positive | ybt S,entB, mrkD, rmpA |
| Kp46 | urine | female | positive | ybt S,entB, mrkD, rmpA |
| Kp53 | urine | female | positive | ybt S,entB, mrkD |
| Kp58 | urine | female | positive | entB, iutA |
| Kp60 | urine | female | positive | entB |
Screening for ESBLs and results hypermucoviscosity K. pneumoniae (hv-KP) strains
The results of screening for the ESBL showed that 62 isolates (88.6%) were ESBL-producing K. pneumoniae isolates. Of the 70 clinical isolates, 10 isolates (14.3%) were positive and 60 isolates (85.7%) were negative for the hv-KP test. Table 2 shows characteristics of hypermucoviscosity clinical isolates.
Virulence genes identification
According to the results of screening virulence genes, entB (n= 57, 81.4%) was the most prevalent among all the clinical isolates, followed by the mrkD (n=46, 65.7 %%), ybtS (n=42, 60%), iutA (n=8, 11.4%), kfu (n=8, 11.4%), rmpA (n=4, 5.7%), magA (n=1, 1.43%), and K2 was not detected in any of the isolates. The presence of the ybtS, entB, mrkD, magA, kfu, iutA, rmpA and K2 genes coding virulence factors was detected among the ESBL and non-ESBL K. pneumoniae isolates (Table 3).
| Total isolates | ESBL | Non-ESBL | P-Value | |
| ybt S | 42(60%) | 38(90.5%) | 4(9.5%) | 0.705 |
| entB | 57(81.4%) | 52(91.2%) | 5(8.8%) | 0.161 |
| mrkD | 46(65.7%) | 43(93.5%) | 3(6.5%) | 0.113 |
| magA | 1(1.4%) | 1(100%) | 0(0%) | 1.000 |
| iutA | 8(11.4%) | 6(75%) | 2(25%) | 0.225 |
| Kfu | 8(11.4%) | 7(87.5%) | 1(12.5 %) | 1.000 |
| rmpA | 4(5.7%) | 4(100%) | 0(0%) | 1.000 |
| K2 | 0(0 %) | 0(0%) | 0(0 %) | 0.231 |
Discussion
Generally, one of the classes of antibiotics used for treating K. pneumoniae is beta-lactams such as cephalosporin 17. However, the presence of ESBL enzymes impairs the performance of these antibiotics 18. The difference in sensitivity and drug resistance in different geographic regions can be associated with different patterns of antibiotic use in different areas 19. In this survey, 88.6% of the clinical isolates were the ESBL producers. Moreover, as shown in the study by Ghasemi, 60% of K. pneumoniae isolates were ESBL producers in Shiraz, Iran 20. Jaskulski et al. in Brazil reported that all K. pneumoniae isolates were ESBL-positive. The prevalence of ESBL-producing clinical isolates is related to different risk factors such as current antibiotic use, resent hospitalization 21.
K. pneumoniae has many virulence factors such as capsular polysaccharide, adhesions and siderophores which contribute to the pathogenicity of these bacteria. Presence of virulence factors in K. pneumoniae is important because they are the most prominent cause of death in patients before starting antibiotic therapy 15. YbtS, entB, mrkD, magA, kfu, iutA, rmpA and K2 genes are among genes that code virulence factors 22. Our study focused on detection of ybtS, entB, mrkD, magA, kfu, iutA, rmpA, and K2 genes in ESBL and non-ESBL producing K. pneumoniae isolates. The important point in this study is that it was the first study to report the presence of virulence genes in K. pneumoniae isolates in Kurdistan Province, Iran. So far, there has been no report of virulence genes in K. pneumoniae on Google Scholar and PubMed. In the present study, entB was determined in 81.43% of the isolates whereas no isolates carried K2 among all the K. pneumoniae isolates taken from the clinical specimens. Nevertheless, entB and K2 were the highest and lowest prevalent virulence factors in the current study. In this investigation, among the 70 isolates collected from clinical specimens such as blood, tracheal, wound, and urine, 10 isolates (14.3%) were hv-KP isolates.
Frequency of rmpA was (n=4, 5.7 %) that all the isolates were ESBL. According to table 2, this is while all the hvKP-isolates had the entB gene and ESBL phenotype. In contrast, in previous studies, such as Yu et al. in Taiwan, the prevalence of hv-KP, rmpA, and magA was reported to be 38%, 48% and 17%, respectively; the result of their study showed that strains carrying rmpA were significantly associated with hv-Kp 23. On the other hand, Nahavandinejad et al. in northern Iran demonstrated that the hv-KP isolates were not restricted to magA 24. MagA was only found in one ESBL isolate that contained the ybtS, entB, mrkD, and kfu genes. In contrast, MagA was much higher than the magA and K2 genes detected in Korea 25 and Taiwan 8. These difference between the prevalence of MagA and K2 could be related to sample type of infection 26. In the majority of those studies, K. pneumoniae was isolated from liver and meninges curtains infections whereas in our study, the isolates were collected from the clinical specimens 28. In a study conducted by Feizabadi et al. in Iran on 89 isolates of K. pneumoniae, 10 (11.2%) isolates belonged to K1 and 13 (14.6%) isolates belonged to K2 serotypes, respectively 27. Amraie et al. in Shahrekord, Iran, reported low frequency of MagA among clinical isolates, which is similar to our results 26. Prior studies suggested that the magA gene can be infrequently seen in K. pneumoniae isolated clinical samples except liver abscesses 2826. Compain et al. in France designed a multiplex PCR for identifying seven virulence factors and K1/K2 capsular serotypes of K. pneumoniae. The multiplex PCR was used on 65 K. pneumoniae isolates between 2004 and 2014, which included 45 clinical isolates identified as hvKP; most isolates (64 /65) were found to possess mrkD 14 which is dissimilar to our results. Unfortunately, there has been no report of virulence genes in ESBL and non ESBL K. pneumoniae.
As a result of these investigations, the presence of virulence genes in ESBL-producing isolates more than clinical isolates of K. pneumoniae lacking ESBL. Our results indicated that there were no statistically significant differences between the ESBL productions and presence of the ybtS, entB, mrkD, magA, kfu, iutA, rmpA, and K2 genes. Moreover, the presence of these genes and variables such as presence of sex, clinical specimen type and hv-KP phenotype between ESBL and non –ESBL K. pneumoniae isolates (0.05< p).
Conclusions
In conclusion, frequency of ESBL-producing K. pneumoniae is increasing now. Detection of virulence factors that positively impact the pathogenicity of K. pneumoniae is of immense importance. The results of the current study showed that entB was the major virulence factor for K. pneumoniae (ESBL and non-ESBL) isolated from the clinical specimens in the hospitals of Kurdistan Province, Iran.
Open Access
This article is distributed under the terms of the Creative Commons Attribution License (CCBY4.0) which permits any use, distribution, and reproduction in any medium, provided the original author(s) and the source are credited.
List of abbreviations
CDDT: Combination disk diffusion test; ESBL: Extended-spectrum betalactamase; LPS: Lipopolysaccharide; MDR: Multi-drug resistant; PCR: Polymerase Chain Reaction; UTI: Urinary tract infection
Ethics approval and consent to participate
The study was approved by Kurdistan University of medical science, Iran. All the members were fully informed of the purpose of the investigation, and were informed.
Competing interests
The authors declare no conflict of interest.
Funding
None
Authors' contributions
PS: Study design, doing experiments, data collection, writing; MKT: Study design, writing, critical review; RR: Supervision, study design, writing, critical review; AA: Data collection, data analysis, critical review; SR: Doing experiments, data collection, data analysis, critical review.
Acknowledgments
This work was retrieved from the thesis of PhD student, Pegah Shakib and, Kurdistan University of Medical Sciences supported this study.
References
-
Podschun
R.,
Ullmann
U..
Klebsiella spp. as nosocomial pathogens: epidemiology, taxonomy, typing methods, and pathogenicity factors. Clinical Microbiology Reviews.
1998;
11
:
589-603
.
-
Fang
C. T.,
Chuang
Y. P.,
Shun
C. T.,
Chang
S. C.,
Wang
J. T..
A novel virulence gene in Klebsiella pneumoniae strains causing primary liver abscess and septic metastatic complications. The Journal of Experimental Medicine.
2004;
199
:
697-705
.
-
M. Azadpour,
J. Nowroozi,
G.R. Goudarzi,
H. Mahmoudvand.
Presence of qacEΔ1 and cepA genes and susceptibility to a hospital biocide in clinical isolates of Klebsiella pneumoniae in Iran. Trop Biomed. .
2015;
32
(1)
:
109-115
.
-
Chander
A.,
Shrestha
C. D..
Prevalence of extended spectrum beta lactamase producing Escherichia coli and Klebsiella pneumoniae urinary isolates in a tertiary care hospital in Kathmandu, Nepal. BMC Research Notes.
2013;
6
:
487
.
-
Drawz
S. M.,
Papp-Wallace
K. M.,
Bonomo
R. A..
New β-lactamase inhibitors: a therapeutic renaissance in an MDR world. Antimicrobial Agents and Chemotherapy.
2014;
58
:
1835-46
.
-
Kil
K. S.,
Darouiche
R. O.,
Hull
R. A.,
Mansouri
M. D.,
Musher
D. M..
Identification of a Klebsiella pneumoniae strain associated with nosocomial urinary tract infection. Journal of Clinical Microbiology.
1997;
35
:
2370-4
.
-
Lery
L. M.,
Frangeul
L.,
Tomas
A.,
Passet
V.,
Almeida
A. S.,
Bialek-Davenet
S..
Comparative analysis of Klebsiella pneumoniae genomes identifies a phospholipase D family protein as a novel virulence factor. BMC Biology.
2014;
12
:
41
.
-
Yeh
K. M.,
Lin
J. C.,
Yin
F. Y.,
Fung
C. P.,
Hung
H. C.,
Siu
L. K..
Revisiting the importance of virulence determinant magA and its surrounding genes in Klebsiella pneumoniae causing pyogenic liver abscesses: exact role in serotype K1 capsule formation. The Journal of Infectious Diseases.
2010;
201
:
1259-67
.
-
Lin
Y. C.,
Lu
M. C.,
Tang
H. L.,
Liu
H. C.,
Chen
C. H.,
Liu
K. S..
Assessment of hypermucoviscosity as a virulence factor for experimental Klebsiella pneumoniae infections: comparative virulence analysis with hypermucoviscosity-negative strain. BMC Microbiology.
2011;
11
:
50
.
-
Lin
J. C.,
Koh
T. H.,
Lee
N.,
Fung
C. P.,
Chang
F. Y.,
Tsai
Y. K..
Genotypes and virulence in serotype K2 Klebsiella pneumoniae from liver abscess and non-infectious carriers in Hong Kong, Singapore and Taiwan. Gut Pathogens.
2014;
6
:
21
.
-
Osman
K. M.,
Hassan
H. M.,
Orabi
A.,
Abdelhafez
A. S..
Phenotypic, antimicrobial susceptibility profile and virulence factors of Klebsiella pneumoniae isolated from buffalo and cow mastitic milk. Pathogens and Global Health.
2014;
108
:
191-9
.
-
Yeh
K. M.,
Kurup
A.,
Siu
L. K.,
Koh
Y. L.,
Fung
C. P.,
Lin
J. C..
Capsular serotype K1 or K2, rather than magA and rmpA, is a major virulence determinant for Klebsiella pneumoniae liver abscess in Singapore and Taiwan. Journal of Clinical Microbiology.
2007;
45
:
466-71
.
-
Enani
M. A.,
El-Khizzi
N. A..
Community acquired Klebsiella pneumoniae, K1 serotype. Invasive liver abscess with bacteremia and endophthalmitis. Saudi Medical Journal.
2012;
33
:
782-6
.
-
Compain
F.,
Babosan
A.,
Brisse
S.,
Genel
N.,
Audo
J.,
Ailloud
F..
Multiplex PCR for detection of seven virulence factors and K1/K2 capsular serotypes of Klebsiella pneumoniae. Journal of Clinical Microbiology.
2014;
52
:
4377-80
.
-
Lawlor
M. S.,
O’connor
C.,
Miller
V. L..
Yersiniabactin is a virulence factor for Klebsiella pneumoniae during pulmonary infection. Infection and Immunity.
2007;
75
:
1463-72
.
-
Beyene
G.,
Tsegaye
W..
Bacterial uropathogens in urinary tract infection and antibiotic susceptibility pattern in jimma university specialized hospital, southwest ethiopia. Ethiopian Journal of Health Sciences.
2011;
21
:
141-6
.
-
Ongut
G.,
Daloglu
A. E.,
Baysan
B. O.,
Daglar
D.,
Ogunc
D.,
Sekercioglu
A. O..
Evaluation of a chromogenic medium for detection of extended-spectrum-beta-lactamase-producing Escherichia coli and Klebsiella pneumoniae strains. Clinical Laboratory.
2014;
60
:
1213-5
.
-
Cassu-Corsi
D.,
Martins
W. M.,
Scheffer
M. C.,
Cayô
R.,
Gales
A. C..
Misidentification of pan drug-resistant Klebsiella pneumoniae clinical isolates as a metallo-β-lactamase producers by the EDTA/DDST test. The Brazilian Journal of Infectious Diseases.
2015;
19
:
102-4
.
-
Ruiz-Garbajosa
P.,
Curiao
T.,
Tato
M.,
Gijón
D.,
Pintado
V.,
Valverde
A..
Multiclonal dispersal of KPC genes following the emergence of non-ST258 KPC-producing Klebsiella pneumoniae clones in Madrid, Spain. The Journal of Antimicrobial Chemotherapy.
2013;
68
:
2487-92
.
-
Ghasemi
Y.,
Archin
T.,
Kargar
M.,
Mohkam
M..
A simple multiplex PCR for assessing prevalence of extended-spectrum β-lactamases producing Klebsiella pneumoniae in Intensive Care Units of a referral hospital in Shiraz, Iran. Asian Pacific Journal of Tropical Medicine.
2013;
6
:
703-8
.
-
Jaskulski
M. R.,
Medeiros
B. C.,
Borges
J. V.,
Zalewsky
R.,
Fonseca
M. E.,
Marinowic
D. R..
Assessment of extended-spectrum β-lactamase, KPC carbapenemase and porin resistance mechanisms in clinical samples of Klebsiella pneumoniae and Enterobacter spp. International Journal of Antimicrobial Agents.
2013;
42
:
76-9
.
-
Brisse
S.,
Fevre
C.,
Passet
V.,
Issenhuth-Jeanjean
S.,
Tournebize
R.,
Diancourt
L..
Virulent clones of Klebsiella pneumoniae: identification and evolutionary scenario based on genomic and phenotypic characterization. PLoS One.
2009;
4
:
e4982
.
-
Yu
W. L.,
Ko
W. C.,
Cheng
K. C.,
Lee
H. C.,
Ke
D. S.,
Lee
C. C..
Association between rmpA and magA genes and clinical syndromes caused by Klebsiella pneumoniae in Taiwan. Clinical Infectious Diseases.
2006;
42
:
1351-8
.
-
Nahavandinejad
M.,
Asadpour
L..
Mucoviscosity Determination and Detection of magA and rmpA Genes in Clinical Isolates of Klebsiella pneumoniae in Northern Iran. Crescent Journal of Medical and Biological Sciences.
2017;
4
:
104-7
.
-
Chung
D. R.,
Lee
H. R.,
Lee
S. S.,
Kim
S. W.,
Chang
H. H.,
Jung
S. I..
Evidence for clonal dissemination of the serotype K1 Klebsiella pneumoniae strain causing invasive liver abscesses in Korea. Journal of Clinical Microbiology.
2008;
46
:
4061-3
.
-
Amraie
H.,
Shakib
P.,
Rouhi
S.,
Bakhshandeh
N.,
Zamanzad
B..
Prevalence assessment of magA gene and antimicrobial susceptibility of Klebsiella pneumoniae isolated from clinical specimens in Shahrekord, Iran. Iranian Journal of Microbiology.
2014;
6
:
311-6
.
-
Feizabadi
M. M.,
Raji
N.,
Delfani
S..
Identification of Klebsiella pneumoniae K1 and K2 capsular types by PCR and quellung test. Jundishapur Journal of Microbiology.
2013;
6
:
e7585
.
-
Fang
F. C.,
Sandler
N.,
Libby
S. J..
Liver abscess caused by magA+ Klebsiella pneumoniae in North America. Journal of Clinical Microbiology.
2005;
43
:
991-2
.
Comments
Downloads
Article Details
Volume & Issue : Vol 5 No 8 (2018)
Page No.: 2581-2589
Published on: 2018-08-24
Citations
Copyrights & License

This work is licensed under a Creative Commons Attribution 4.0 International License.
Search Panel
Pubmed
Google Scholar
Pubmed
Google Scholar
Pubmed
Google Scholar
Pubmed
Google Scholar
Pubmed
Search for this article in:
Google Scholar
Researchgate
- HTML viewed - 8803 times
- Download PDF downloaded - 3283 times
- View Article downloaded - 0 times
 Biomedpress
Biomedpress


